RESEARCH HIGHLIGHTS
Our research highlights serve as a collection of feature articles detailing recent scientific achievements on GCS HPC resources.
OneProt Initiative Pioneers a Multi-Modal Foundation Model for Protein Research
Research Highlight –
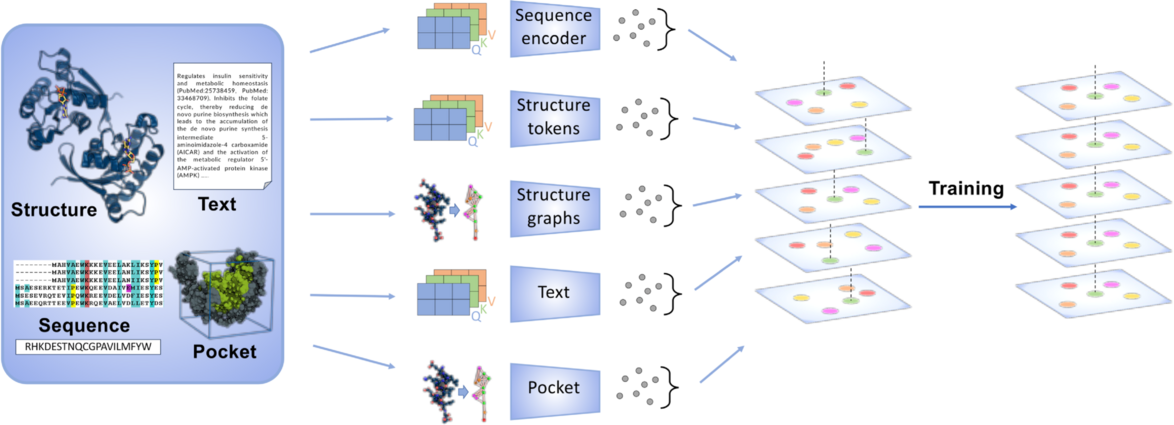
The OneProt model aligns multiple modalities, including primary protein sequence, 3D protein structure, protein binding sites, and text annotations. Each modality is processed by its respective encoder, generating embeddings aligned in a shared latent space, facilitating cross-modal learning and integration. Image credit: Alina Bazarova, JSC.
Computational biologists and public health researchers rely on modeling and simulation to design new medications and to better understand how to deliver those drugs where they belong in the body. From speeding up vaccine development to gaining a deeper understanding of how certain viruses invade our body, simulations using high-performance computing (HPC) have become essential tools for advancing healthcare.
While traditional modeling and simulation methods remain important pillars for research into protein structure and function, rapidly advancing artificial intelligence (AI) methods offer powerful new tools to speed up research. “With AI, we are now capable of processing even larger amounts of information and biological detail on HPC systems, which has had a great impact on this kind of research,” said Dr. Alina Bazarova, AI consultant at the Jülich Supercomputing Centre (JSC). “Now, as more diverse AI architectures are being developed, we are able to take more protein properties into account, and we can train more complex models for longer amounts of time.”
Bazarova is JSC’s institutional contact for the OneProt initiative. OneProt, which includes collaborators from Helmholtz AI, the Technical University of Munich, and other research groups at Forschungszentrum Jülich, focused on producing a multi-modal foundation model for protein research. This essentially means that the model is trained to process disparate datasets related to proteins and can be adapted by various research teams to focus on their specific research questions. The initial model was revealed in the team’s recent paper in PLOS Computational Biology and sets the stage for the next phase of the team’s work—OneProtGPT, which integrates large language models (LLMs) into this powerful AI tool.
Multi-modal foundation models support larger research communities
The original motivation for OneProt began with researchers planning to focus on creating embeddings — essentially small numerical representations of proteins that allow protein variants to be studied by machine learning algorithms — for different protein mutations. During the development of the system, the researchers realized that training an AI system for this task could be readily extrapolated into other relevant research tasks. Therefore, the development team devised an easy-to-use fine-tuning process that only requires one to two additional layers in the model’s neural network and can be adapted to any task without retraining. Additionally, to make OneProt framework successful as a foundation model, the team designed it so researchers could easily incorporate new kinds of data during pre-training.
Accordingly, the model is not only capable of supporting research into protein mutations, but also can be adapted for studying the molecular structure of proteins and gene ontology—essentially a classification system for gene function—among other research questions. OneProt was trained using massive amounts of data related to proteins’ 3D structure, amino acid sequences, and binding site details, and the model has performed especially well at tasks predicting enzyme function and analyzing protein binding sites. The team attributes some of that success—as well as the model’s relatively lean training footprint—to training using multiple modalities, meaning essentially using various different types of data related to protein function and behavior. “Considering the improvements to AI models, we designed OneProt so that it could function as somewhat more of a black-box technology,” Bazarova said. “Rather than using an HPC system to take into account every possible interaction between protein molecules, we created some functions that help us get the desired outputs by allowing the AI model itself to infer some of these comparisons, as it is better at disentangling this data than we are on our own.”
OneProtGPT promises to further streamline foundation model via large-language models
Having demonstrated success with OneProt, the team applied for time on JSC’s newest flagship supercomputer—JUPITER, Europe’s first exascale system—as part of the Gauss AI Compute Competition. The program, organized by the Gauss Centre for Supercomputing, focuses on advancing next-generation generative AI models that can benefit diverse research communities. The work is funded by the German Federal Ministry of Research, Technology, and Space, the North Rhine-Westphalia Ministry of Culture and Science, the Baden-Württemberg Ministry of Science, Research and Arts, and the Bavarian State Ministry of Science and the Arts.
“OneProt was trained on the JUWELS Booster system, which uses NVIDIA A100 GPUs, and we are now moving to JUPITER, which offers GH200 Grace Hopper superchips,” said Javad Kasravi, HPC AI research engineer at JSC and collaborator on the initiative. “The GH200 configuration combines larger memory, higher compute capacity, and faster data transfer. With roughly three times the GPU memory compared with the A100, an NVIDIA Grace CPU, and a high-speed link between the CPU and GPU, we can scale training more efficiently—helping us iterate faster and improve model accuracy in less time.”
With access to JUPITER, the OneProt team is actively improving the model by integrating protein data with large language models. By integrating LLMs into OneProt, researchers could generate more detailed protein descriptions when using various data sources. “For now, we have fine-tuned OneProtGPT using protein sequence data, but we are working on the ability to replace this sequence data by other data modalities, which would greatly expand the model’s flexibility for a variety of applications,” Bazarova said.
-Eric Gedenk
Funding for JUWELS was provided by the Ministry of Culture and Research of the State of North Rhine-Westphalia and the German Federal Ministry of Research, Technology and Space through the Gauss Centre for Supercomputing (GCS).
Related Publication: Flöge K, Udayakumar S, Sommer J, Piraud M, Kesselheim S, et al. (2025). “OneProt: Towards multi-modal protein foundation models via latent space alignment of sequence, structure, binding sites and text encoders,” PLOS Computational Biology 21(11). DOI: https://doi.org/10.1371/journal.pcbi.1013679