LIFE SCIENCES
A Molecular View on How Bacteria Fight Toothpaste Fluoride
Principal Investigator:
Dr. Mercedes Alfonso-Prieto & Dr. Emiliano Ippoliti
Affiliation:
Forschungszentrum Jülich GmbH, INM-9 Computational Biomedicine, Jülich, Germany
Local Project ID:
fluc-gs
HPC Platform used:
JUWELS at JSC
Date published:
Abstract/Teaser
Researchers from Jülich and University of Milan Bicocca investigated how microorganisms protect themselves from the toxic effects of fluoride. They focused on a protein called Fluc, which creates a tunnel in the membrane to pump fluoride out from bacterial cells. One of the tunnel stations for fluoride contains a protein amino acid, glutamate, which can be negatively charged, as fluoride is. However, two close negative charges would repel each other and bring Fluc to a halt. Such jam could be resolved by either lifting the glutamate barrier out of the way or paying the toll of protonating the glutamate. Preliminary results from multiscale molecular dynamics simulations suggest that both options are feasible.
Project
You might have heard of fluoride as an ingredient of toothpaste and mouthwash, but did you know that fluorine is the 13th most abundant element in the Earth crust? It is present everywhere, from water to soil, posing a problem for microorganisms, for which fluoride is toxic. The negatively charged fluoride (F–) can disguise itself by gaining a proton (H+) and becoming the neutral HF (hydrofluoric acid), which can easily cross the membranes surrounding bacterial cells and reach their interior. Once inside bacteria, fluoride loses the proton and accumulates in the bacterial cytoplasm, causing toxicity and eventually cell death. Hence, bacteria have developed strategies to combat fluoride. One of those are fluoride channels (Flucs), proteins that create a tunnel in the bacterial membrane to pump fluoride out and thus avoid toxicity. Based on experimental data [1] and previous simulations [2-3], it was proposed that this tunnel has three stations or fluoride binding sites (F0, F1 and F2) and that fluoride stops in each of these stops until another incoming fluoride pushes it to move to the next station. F2 is the last station in this tunnel and contains a protein amino acid, glutamate, that is conserved across bacteria. Nature choosing to place a glutamate in F2 is somewhat surprising, as both fluoride and glutamate are in principle negatively charged and thus would repel each other when they are close, stopping fluoride traffic through the tunnel. Two hypotheses have been proposed to solve this apparent electrostatic jam: either lifting the glutamate[-] barrier out of the way or paying the toll of adding a proton or protonating it (glutamate[H]).
Molecular dynamics (MD) is an excellent computational tool for observing fluoride ions as they go through the tunnel, akin to a computational microscope. In this project, we employed two different focuses or resolutions to study how fluoride goes through the F2 station. First, classical (molecular mechanics-based) MD simulations gave us an insight into the movement of the glutamate barrier. Second, we adjusted our focus to be able to watch the proton moving from/to glutamate and fluoride. This was accomplished by using a multiscale approach, called quantum mechanics/molecular mechanics (QM/MM), which allows simulating one part of the system (that is the F2 site) at a sharper resolution than the rest (including the remainder of the tunnel, the bacterial membrane and the surrounding solution).
These simulations were only possible thanks to the computational resources granted on the JUWELS supercomputer at the Jülich Supercomputing Centre. In order to mimic the physiological conditions, the Fluc channel must be embedded in a membrane and solvated with water and ions, resulting in a system size of approximately 100,000 atoms. Moreover, we tested twelve different local conditions at the F2 station and ran three replicates for each, for a total of 36 simulations. Each of these classical simulations can be considered a movie that is 500 ns long and composed of 250,000 frames. Therefore, we needed JUWELS to perform 36 times 250,000 or nine million calculations! Next, we selected representative snapshots of these classical simulations to carry out QM/MM simulations. Increasing the level of resolution for the F2 site comes at a significantly higher computational cost. Therefore, we needed a program that was able to utilize as efficiently as possible the supercomputing resources. For this, we relied on the MiMiC framework for multiscale modeling in computational chemistry [4].
This is not the end of the story: We are still trying to understand how the glutamate barrier lifting and the protonation toll fee can act together to help fluoride flow through the F2 station. Such knowledge would help us understand how Fluc channels can function as bidirectional tunnels, allowing fluoride traffic in both directions through a single lane.
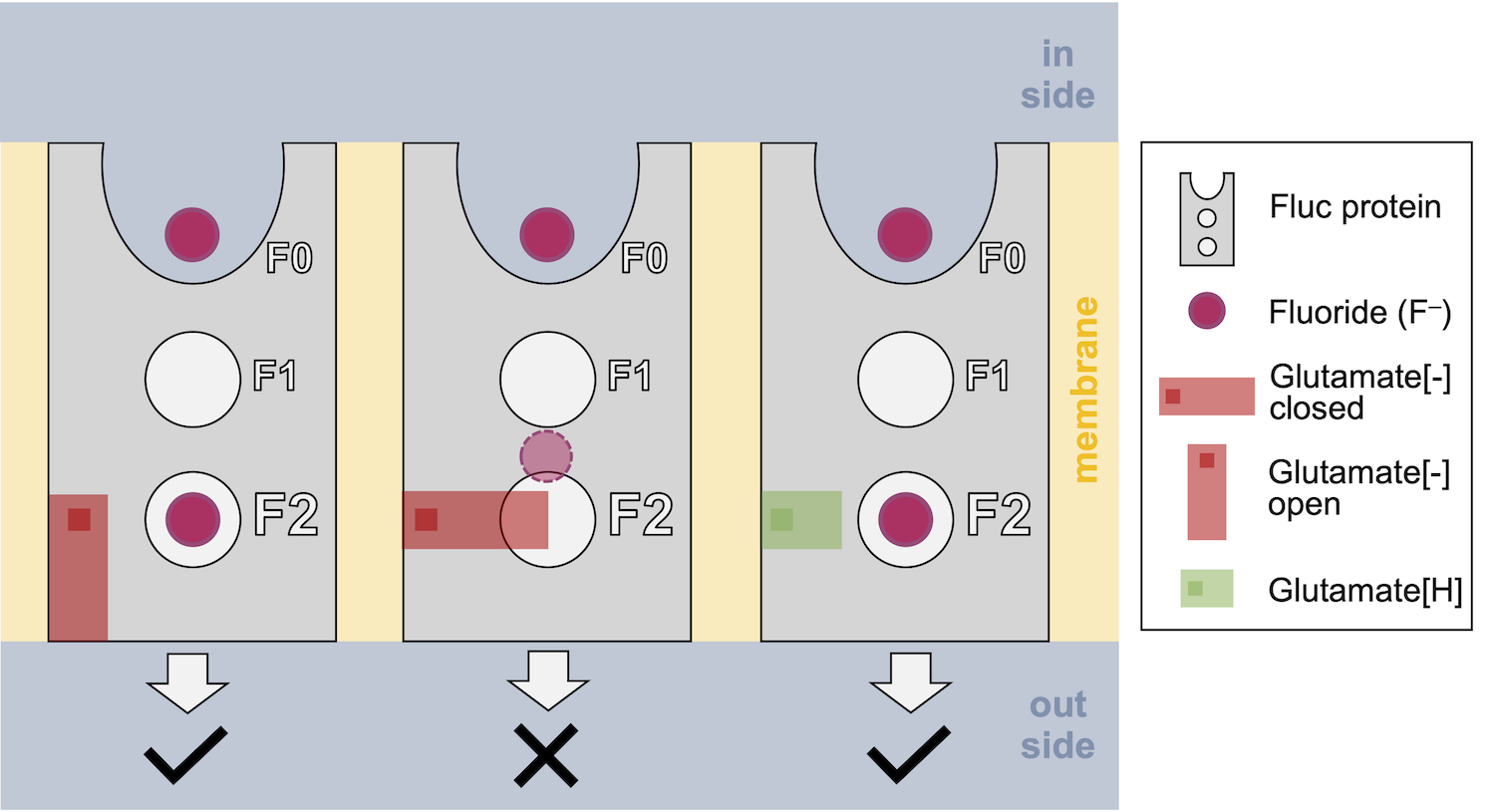
The fluoride channel Fluc (gray) pumps out fluoride ions (F–, in purple) from the bacterial inside to the outside to prevent toxicity. The Fluc channel has three stations, F0, F1 and F2, with the last one regulating fluoride traffic by using a glutamate barrier. When the barrier is closed (central image, glutamate[-] in red), fluoride is stopped. Fluc can restore traffic using two alternative solutions, either by lifting the barrier (left image, glutamate[-] in red) or paying the toll of adding a proton (H+) to the glutamate (right image, glutamate[H] in green).
References
[1] McIlwain, B. C et al. “The fluoride permeation pathway and anion recognition in Fluc family fluoride channels.” eLife (2021) 10: e69482.
[2] Yue, Z. et al. “Ion permeation, selectivity, and electronic polarization in fluoride channels.” Biophysical Journal (2022) 121(7): 1336-1347.
[3] Zhang, J. et al. “Fluoride permeation mechanism of the Fluc channel in liposomes revealed by solid-state NMR.” Science Advances (2023) 9:eadg9709.
[4] Raghavan B. et al. “Drug design in the exascale era: a perspective from massively parallel QM/MM simulations.” Journal of Chemical Information and Modeling (2023) 63(12): 3647–3658.