LIFE SCIENCES
Enabling Genomic-Assisted Breeding of Cattle and Chickens in Developing Countries
Principal Investigator:
Prof. Dr. Laurent Frantz
Affiliation:
Ludwig Maximilians Universität München, Tierärztliche Fakultät, Institut für Paläoanatomie und Geschichte der Tiermedizin, Munich, Germany
Local Project ID:
pn29qe
HPC Platform used:
SuperMUC-NG at LRZ
Date published:
Introduction
Intensified production has led to remarkable gains in chicken production (5x), growth rate (3x), and milk yield (2x). However, this relies on narrowed genetic diversity in primarily European commercial breeds, increasing their vulnerability to future biotic (disease) and abiotic (climate) stresses. Local breeds, critical for adaptation, are experiencing alarming declines (34% endangered, 5% extinct in 15 years) [1]. This is driven by the adoption of high-yielding European breeds, particularly in Africa. While initially productive, these breeds are often poorly adapted to local environments and lack disease resistance. Our research leverages genomics to understand the impact of industrialization and globalization on livestock genetics. We analyze thousands of diverse genomes to reconstruct history, characterize adaptations, and assess threats like inbreeding. Additionally, ancient DNA analysis reveals how livestock species have evolved, informing sustainable management strategies that balance productivity with genetic diversity and adaptability. This approach, combined with our large computationally intensive genomic dataset, offers the potential to develop genomic-based breeding tools for underutilized breeds. These breeds, while lower-performing than their highly selected counterparts, possess local adaptations that enhance their long-term sustainability. While European breeds offer short-term benefits for non-European smallholder farmers, like in Africa, their vulnerability to drought and local diseases makes them risky investments in the long term, especially considering the vital role livestock plays as savings in communities as in Sub-Saharan Africa [2]. Therefore, we aim to pave the way for genomic-assisted breeding (genomic selection) of local breeds. This technology, successfully used in Europe to improve productivity while limiting inbreeding, requires high-quality reference imputation panels representative of complex livestock (chicken and cattle) ancestries. This will enable the deployment of genomic breeding tools, promoting sustainable livestock management in developing regions.
Results and Methods
This project has leveraged a state-of-the-art pipeline deployed on the SuperMUC-NG to assemble large genomic datasets for chickens and cattle.
Chicken Dataset [3]:
- 4,392 genomes encompassing domesticated chickens, red junglefowl subspecies, and congeneric Gallus species.
- Diverse geographical origins representing commercial birds, experimental lines, and local breeds.
- ~50 ancient genomes reveal past genetic diversity and inform domestication history.
Cattle Dataset:
- 128 newly generated genomes from 32 diverse African populations.
- Integrated with publicly available data for a total of 3,527 genomes.
These comprehensive datasets enable deeper understanding of genetic diversity, adaptation, and evolutionary history in both species, informing sustainable management strategies.
Data processing pipeline
To ensure uniformity across all samples, the entire dataset was subjected to a state-of-the-art genomic pipeline on the SuperMUC-NG computing cluster. The processing of these genomes required ~25 M core hours.
Imputation pipeline
Large-scale genomic datasets, processed on the SuperMUC-NG, were merged into reference panels containing tens of millions of genetic variants. These panels enable the deployment of low-coverage genome sequencing with imputation, a cost-effective tool for genomic-assisted breeding (genomic selection) in developing countries. Briefly, imputation computationally predicts missing genotypes based on known variants in reference panels.
Evaluating Imputation Accuracy
We assessed the performance of our reference panel for imputation. Traditionally, reliable genome data requires 10x coverage, exceeding €200 per sample. Using our imputation pipeline and reference panel, we achieved similar quality with 0.5x coverage. This will reduce the cost of generating genome data to less than €20 per sample. This significant cost reduction empowers animal breeders in developing countries to generate data for improving local breeds. This opens avenues for:
- Sustainable breeding: Selecting animals with desired traits based on their genetic makeup, leading to improved productivity and adaptation.
- Preserving diversity: Characterizing and conserving genetic diversity within local breeds, ensuring their resilience to future challenges.
- Empowering local communities: Enabling farmers to make informed breeding decisions and enhance their livelihoods.
By facilitating cost-effective genomic data generation, this approach holds immense potential for advancing sustainable livestock management and empowering developing communities.
Ongoing Research / Outlook
African cattle genomic data
A manuscript detailing our research on imputation accuracy in African cattle is currently under preparation (Figure 1). Additionally, we have generated over 1,000 low-coverage genomes from diverse African cattle populations, which are currently undergoing analysis in conjunction with ancient genomic data. This combined approach aims to elucidate how African cattle have evolved to thrive in a wide range of ecological contexts.
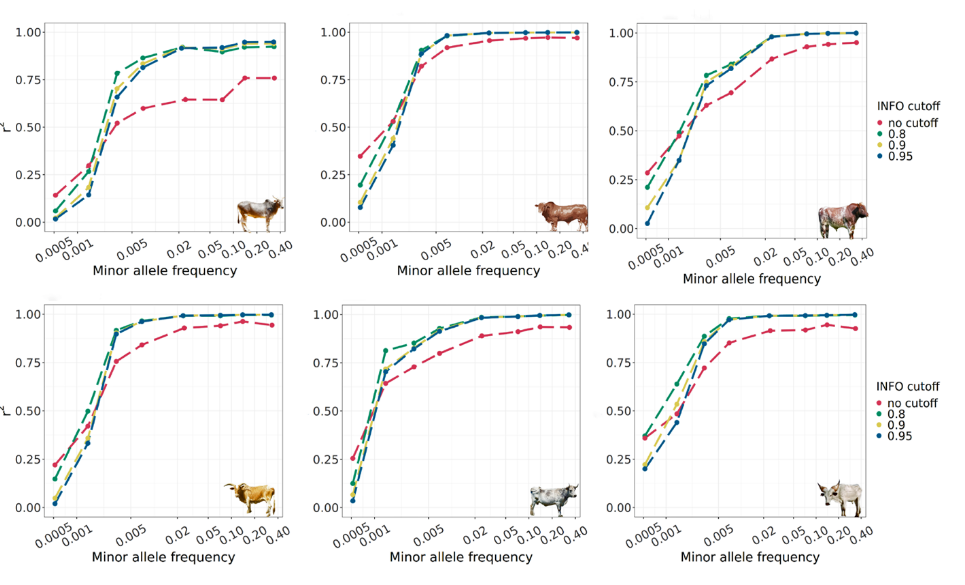
Figure 1: Accuracy of imputation in African cattle breed of 0.5x coverage. The y-axis represents r², with 0 indicating poor imputation quality compared to a 20x coverage genome (≈350) and 1 indicating equivalent quality. The x-axis displays allele frequency, ranging from 0.001 to 0.5. The findings demonstrate that our imputation pipeline can elevate the quality of a 0.5x genome to match that of a 20x genome even for rare mutation (found in 0.2% of cattle individuals). The analyzed breeds encompass diverse geographic origins and genetic backgrounds: Hariana (South Asia), Afrikaner (South Africa), Shorthorn (Europe), Kilimanjaro Zebu (East Africa), Creole (South America), and Kuri (West Africa).
Chicken genomic data
Our research efforts in chickens parallel those undertaken for African cattle. We are currently: 1) Developing an imputation panel. This panel will enable us to assess the effectiveness of imputation in reducing genotyping costs for chickens, paving the way for more affordable genomic studies in this species. 2) Preparing a large-scale chicken genomics paper. This paper, incorporating ancient genomic data, will investigate how chickens, as tropical birds, have adapted to the diverse environments they inhabit today. These findings will contribute to the development of improved breeding strategies and conservation efforts for this vital livestock species.
References and Links
[1] FAO (2013) FAOSTAT database collections. Food and Agriculture Organization of the United Nations.
[2] Menjo D, Bebe B, Okeyo A, Ojango J (2009) Survival of Holstein-Friesian heifers on commercial dairy farms in Kenya. Appl Anim Husb Rural 2:14-17.
[3] Fiddaman SR, Klopp C, Charles M, et al (2023) Chicken Genomic Diversity consortium: large-scale genomics to unravel the origins and adaptations of chickens. Cytogenet Genome Res.